Biological Processes and Interactions
Changes in Molecular Genetics
MINDS ON
Some Key Terms
Look for these terms in this Activity. You should practice using them as you compose your responses to the upcoming tasks.
Realize, though, that this list is not exhaustive so you should look for opportunities to include other relevant terminology related to molecular genetics.
| mutation | gene expression | silent |
| missense | nonsense | frameshift |
| deleterious | inducer | repressor |
| operator | operon | epigenetics |
| methylation | acetylation |
Cell Signaling
Cell-to-cell communication includes hormones, antigens and neurotransmitters. They are made of either hydrophobic steroids or hydrophilic peptides. Watch the video below.
When transcription and translation of a gene is started we say that the gene is being expressed. In this way we can say that certain cell signals alter gene expression.
Genes play an important role in how cells respond to their environment. They are affected by cell-to-cell communication. Interestingly, gene expression can also be changed because of environmental conditions. Human skin colour is an interesting example of this kind of response. See how the pigments of our skin are influenced by both genetics and the environment in this video.
 How does understanding change?
How does understanding change?
Understanding 29. Adaptations help organisms better survive in their environment. Why might darker toned skin be a more common adaptation in countries closer to the equator? How might future changes to the environment lead to further changes in gene expression of skin pigments?
Protein Shapes
Look back at your graphic organizer from Unit 1, Activity 3 on Proteins and your fishbone diagram from Unit 2, Activity 2 on Enzymes to see how the shape of proteins are important for their functions. Amino acids differ from each other with their R-group. In addition to its length we also saw how some side chains are hydrophobic and some are hydrophilic. Among the hydrophilic amino acids, R-groups can be uncharged, positively charged, or negatively charged.
When Evolutionary Biologists look at genetic differences among species they compare the amino acid sequences of a protein that is homologous. (definition:having similar structure in different organisms with a common ancestor.) In this way small changes on the evolutionary level can be used to assemble an evolutionary tree.
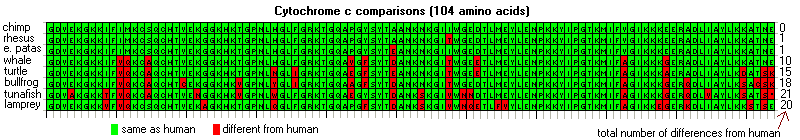
by NMSR
Species with fewer differences in amino acid sequence are more closely related than those with more differences. In the figure above we can see that monkeys like chimpanzees, Rhesus and E. Patas have fewer differences in their cytochrome c proteins compared to humans. This makes sense given that primates are more closely related to humans than other mammals or other chordates. Still, for all species that use an electron transport chain to produce ATP, cytochrome c must be able to function in a homologous way. The question then is, shouldn’t all cytochrome c proteins have the same amino acid sequence in their primary structure?
The answer to this is “perhaps”. We have seen the importance of shape. Indeed, for an important protein involved in energy metabolism any significant changes in the amino acid sequence could be lethal. However, if a substitution of one amino acids for one with similar properties occurs there is a chance that the protein could still fold into its proper tertiary structure. We can see this where at position #11 hydrophobic I (isoleucine) is substituted for hydrophobic V (valine), or at position #12 where hydrophilic M (methionine) is substituted for hydrophobic Q (glutamine) in the non-primate animals.
 How do small steps lead to change?
How do small steps lead to change?
Changes 20. Use the codon table to identify how many different nucleotide substitutions are needed to change the methionine codon to a glutamine codon. How about to change an isoleucine codon to a valine codon?
ACTION
Changing Genes
When we think of the word mutation often the first thing that comes to mind is a monster in a summer blockbuster movie. From a genetic perspective, though, a mutation is just a change in the genetic information of a cell. They can be somatic mutations that happen to our somatic cells or germline mutations that occur in gametes (definition: a cell containing only one copy of chromosome and is used in sexual reproduction.)
 Canadian Contributions
Canadian Contributions
In Grade 11 you learned about changes to chromosomes, like duplications, deletions and insertions. These large-scale mutations were once considered relatively rare. Research by Stephen Scherer showed that certain sections of our genome are copied, sometimes even dozens of times. Some of these duplications are small two- and three-nucleotide repeats while others copy entire genes.
For example the gene for salivary amylase in humans is copied up to nine times in people. Interestingly, the number of copies correlates to the traditional diet of our ancestors. People with ancestry where large amounts of starch are consumed have more copies of the amylase gene. These people also have a higher level of amylase gene expression and produce saliva that is more concentrated in amylase.
Some duplications are also connected to diseases. Others have no effect as they occur in non-coding sections of our genome. Combined, these differences account for the 0.01 % difference in genetic code among each of us. Scherer’s work was also used as part of the Human Genome Project that mapped out the sequence of the human genome.
We are already familiar with many common mutations in genes. Some have a harmful, or deleterious, effect while others are neutral. Variation among individuals in a population is created by these sorts of mutations. Explore some of these examples from Genetic Science Learning Center. Practice good Initiative skills (definition:I can approach new tasks with a positive attitude.) and look to see how the DNA is changed. Also, look to see how mutations in one species can have similar effects in mutations in homologous genes.
 Why is shape important in Biology?
Why is shape important in Biology?
Shape 20. What matters more to how harmful a mutation is to an individual: the size of the mutation, the type of mutation, or the shape of the protein produced? Justify your response with specific examples of mutations.
Single nucleotide mutations, also called point mutations, can be caused by internal or external factors. External factors include radiation, like radioactivity, X-rays or UV light, and chemicals called mutagens, like tar in cigarettes or polycyclic aromatic hydrocarbons from burned fatty meat. Point mutations are sometimes called single nucleotide polymorphisms, particularly for point mutations that have no deleterious effect on a person’s phenotype.
Internal factors involve mistakes by the enzymes responsible for DNA replication. DNA polymerase III has an error rate of inserting a wrong nucleotide once in about 10,000 nucleotides. This causes the double helix to bulge out because the bases can’t properly hydrogen bond there. Recall that towards the end of replication DNA polymerase I proofreads the daughter strands for errors like this and fixes it by putting in the correct nucleotide. Every once in awhile, though, an error stays and is passed to daughter cells after the next round of cell division. And if that error occurs when gametes are formed it might be passed on to the next generation.
Point mutations include single nucleotide deletions, insertions, and substitutions. The effects of these type of points mutations depends on where in the gene they occur. Genes are made of introns and exons. Exon sections combine to include a critical portion such as the active site in enzymes. Amino acids with similar R-groups may not significantly change the shape of the protein. Even within the codons, mutations in different nucleotides in each triplet can have different effects. Watch the following video to see how the structure of genes determines the effect of different point mutations.
In the following interactive, explore gene mutations.
Mutations
 Why is shape important in Biology?
Why is shape important in Biology?
Shape 21. The cause of sickle cell anemia is a point mutation that changes a glutamic acid for a valine amino acid. Why does changing the polarity of this amino acid change the shape of hemoglobin in this disease?
Let’s explore this further by simulating some mutations in the sequence of DNA you made in the previous Activity. This time the simulation will transcribe and translate the sequences for you. Amino acids in the polypeptide chain are coloured to show hydrophobic ones in brown and hydrophilic ones in green.

Controlling Gene Expression
One of the puzzles that Biologists needed to work out was how specialized cells become differentiated. Do specialized cells receive a different set of genetic instructions during specialization from stem cells? Or do specialized cells just read different instructions from the whole genome? Chromosomal studies of different cell types shows that each somatic cell type has a complete set of 46 diploid chromosomes.
Take the example of a firefly’s ability to produce flashes of light. Each somatic cell in the firefly has a full set of genetic instructions in its chromosomes. However, only specialized cells in its abdomen are able to make luciferase, the enzyme responsible for the flashes. Look to see how gene expression is controlled in fireflies in this video from Genetic Science Learning Center.

The way that cells turn off and on genes happens in different ways. In this section we will examine operons, transcription factors, and epigenetics.
 Controlling Gene Expression
Controlling Gene Expression
As you explore these examples think about a graphic organizer you can use to summarize important details of gene expression.
Your summary should include these details and concepts:
- what substance increases or decreases gene expression,
- whether gene expression increases or decreases in response to the substance,
- how RNA polymerase activity changes.
Many prokaryotic genes are organized in groups called operons that are related to a metabolic pathway or cellular processes. The genes in the operon can be induced (turned on) or repressed (turned off) depending on how the genes are used by cells at a given time. For example, E. coli bacteria are able to synthesize the amino acid tryptophan. This anabolic pathway is the product of genes in the trp operon. In order to be energy efficient the amount of tryptophan a cell needs to synthesize is controlled through a negative feedback loop.
We can describe this control of gene expression in a feedback diagram. The expression of the genes in the trp operon is repressed by high tryptophan levels. In this case, tryptophan binds to a repressor molecule and the two molecules together act to prevent transcription of the operon. For this reason tryptophan is called a corepressor.
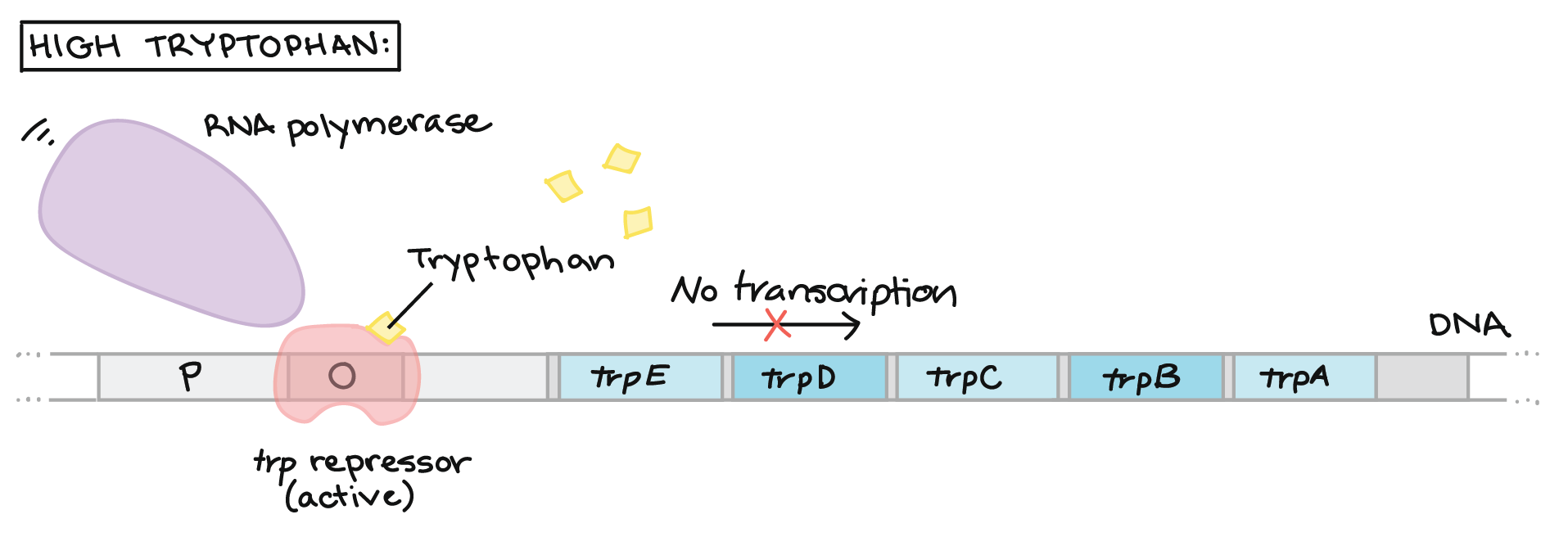
by Khan Academy
Another way to describe control of gene expression is by making a decision tree. Here, the two choices are clearly shown as options and a series of steps branch out from each choice. We can see a decision tree for another operon in E. coli: the lac operon. This catabolic pathway allows cells to metabolize lactose disaccharides. In order to be energy efficient the enzymes needed for lactose metabolism are only expressed when a cell encounters lactose in its environment.
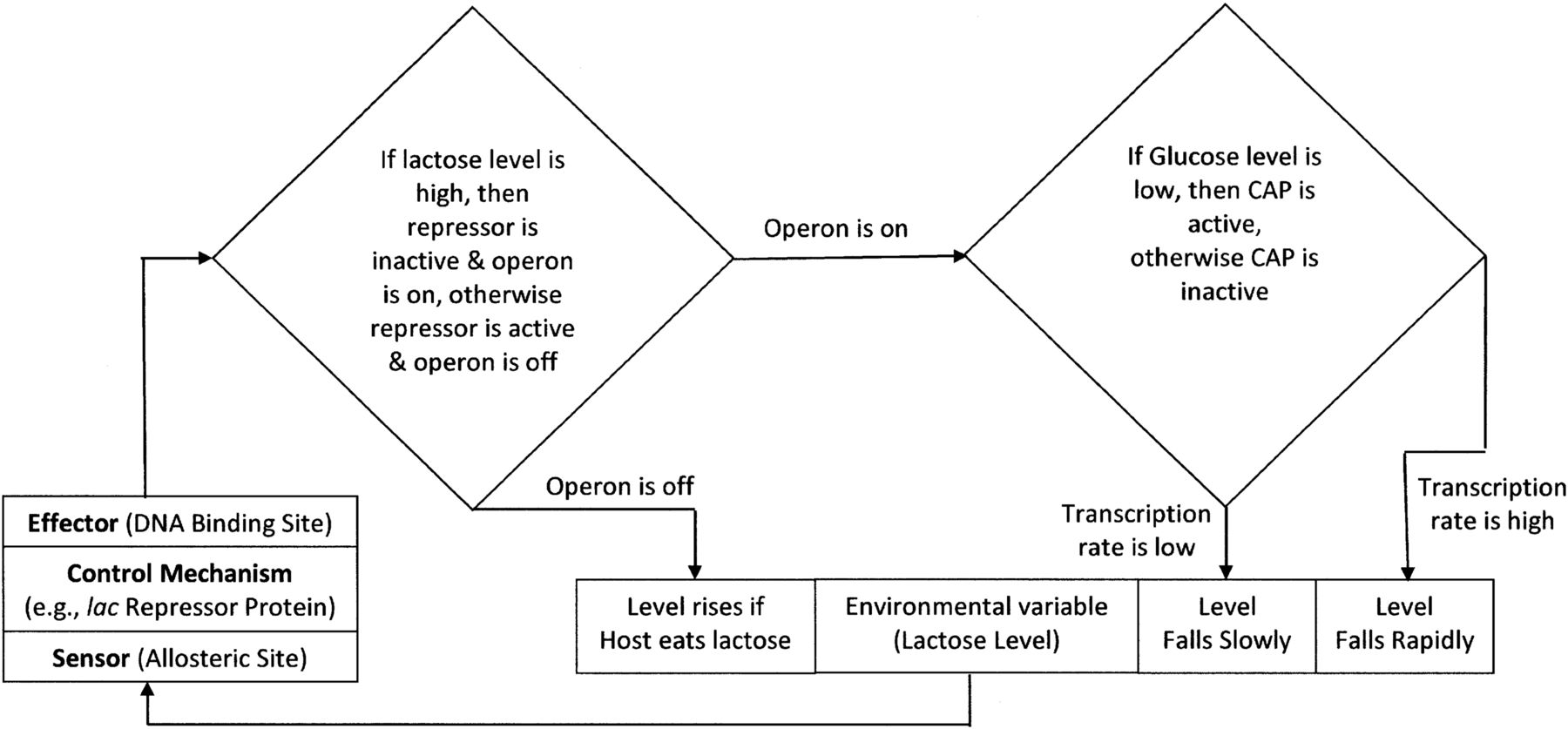
by Robert A. Cooper
The expression of the genes in the lac operon is induced by the presence of lactose. Lactose binds to a repressor and the two molecules together can no longer inhibit RNA polymerase from transcribing the operon. For this reason lactose is called an inducer.
Both types of operon involve a protein molecule called a repressor that prevents RNA polymerase from transcribing. They bind to a section of DNA called the operator that’s between the promoter and the first gene in the operon. Explore these two types of operons further in this interactive to see how changes in the shape of repressors act in both inducible and repressible expressions.
 How do small steps lead to changes?
How do small steps lead to changes?
Changes 21. Compare the effect of lactose and tryptophan on their repressors. How do inducers and corepressors affect gene expression?
In eukaryotes gene expression is more varied. Similar to repressors in operons, transcription factors can alter gene expression. Transcription factors also include activators or enhancers that have positive effects on gene expression. Transcription factors also bind to regions of DNA to help or inhibit RNA polymerase. These regions have different names, such as regulatory region and switch. To see how transcription factors work, first get familiar with the anatomy of a gene by exploring the interactive. Then watch the following video to learn about activators and repressors of gene expression.

Two well studied examples of controlling gene expression in eukaryotes are the circadian rhythm (definition:the daily rhythmic activity cycle, based on 24-hour intervals, that is exhibited by many organisms.) in fruit flies and adult onset lactose intolerance. As you explore these examples further, look for similarities and differences in how transcription is changed.


Eukaryotic genes can also be changed at the chromosomal level. Recall that eukaryotic chromosomes are made of DNA wrapped around histones. When chromatin fibers are opened up then the DNA can be read. Here, two modifications can affect gene expression: methylation (definition:a biochemical reaction that involves the addition of a CH3- functional group.)of nucleotides and modification of histone shape through acetylation. (definition:a biochemical reaction that involves the addition of a CH3CO- functional group.)
Interestingly, these changes are influenced by environmental factors, like nutrition and stress, and can lead to changes in phenotype that can be inherited.
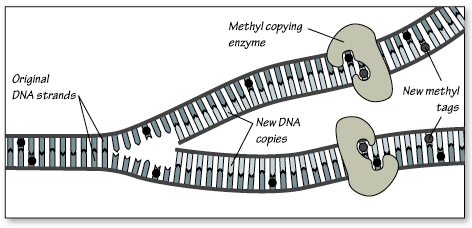
by Genetic Science Learning Center
Because these changes are not part of the DNA code they are called epigenetic changes.
 Why is shape important in Biology?
Why is shape important in Biology?
Shape 22. Methylation and acetylation affect how tight or loose chromatin fibers are. Explain why more compact chromosomes make it more difficult for enzymes involved in the Central Dogma to read DNA? How might this lower levels of gene expression?
Regulatory Genes
Significant research has recently helped to further scientific understanding of how gene expression can be changing in specialized cell types. One puzzling question that has recently been answered had to do with the production of proteins in certain unrelated organs. For example, animals express the pitx1 gene in their jaw, pituitary and pelvis. Mutations in this gene only affected these parts of the body. So, what is the connection between these body parts and this gene?
The answer has to do with general instructions instead of specific instructions. The Pitx1 protein is involved in building structures in the body. The type of structure it helps to build depends on which cell type it’s in. Explore how changes in regulatory genes and the switches that control them in this interactive from Genetic Science Learning Center.
The most famous regulatory genes are called the homeobox, or hox, genes. Watch this video to learn more about these interesting genes.
CONSOLIDATION
Summary
This Activity explored the big idea that gene expression can change. Specifically,
- changes in the sequence of DNA can have beneficial, neutral or deleterious effects;
- transcription can be enhanced or inhibited by changes in a cell’s environment;
- changes in chromosome structure can also change gene expression.
Decisions in Gene Expression
In this Activity we have explored some of the ways that gene expression can change. All of these methods involve natural changes in DNA and chromosomes, even if they are caused by human activity. Research about the role of DNA in the past 30 years has focused on these changes. Increased or decreased gene expression is now being linked to a variety of health outcomes, like diseases, wellbeing, and aging.
The role of methylation seems to play an important role in how cells use DNA. It has been linked to:
- aging,
- cancer,
- embryonic development,
- amount of exercise,
- atherosclerosis,
- the immune system (such as B-cells),
- memory.
 How does understanding change?
How does understanding change?
Choose one of the health outcomes related to methylation listed above. Conduct research to find out whether methylation is increased or decreased. How does the cause of the change in methylation explain its effect on our health? Use specific details and vocabulary about controlling gene expression in your response.