Choices In Biology
Molecular Genetics: Combining Genes in New Ways
MINDS ON
Partial List of Related Materials
In this Activity you will be combining details and concepts from several parts of the course you have already completed. You should refer to them as you compose your responses to the upcoming tasks.
Realize, though, that this list is not exhaustive so you should look for opportunities to include other relevant details related to choices in Biology.
Your answers to: “Why is shape important in Biology?”
- Unit 1, Activity 5 on Nucleic Acids;
- Unit 1, Activity 6 on Cellular Structures;
- Unit 2, Activity 2 on Enzymes;
- Unit 2, Activity 3 on Cellular Transport;
- Unit 2, Activity 9 on Processes in the Central Dogma;
- Unit 2, Activity 10 on Changes in Molecular Genetics
will help your understanding in this activity.
Complementary Base Pairing
Each of the main enzymes involved in the processes of the Central Dogma bind to specific sequences on nucleic acids. We have seen how helicase unwinds DNA at the origin, DNA polymerase III binds to sections of RNA base paired to DNA, RNA polymerase copies DNA at the promoter, and ribosomes scan mRNA molecules from the 5’ cap in search of a start codon. These enzymes “know” how to use the genetic code because of complementary base pairing.
When Biologists began uncovering the genetic code in today’s codon table they were amazed at the universality of it. Most species use the same codon to code for the same amino acid. An interesting question that naturally followed this discovery was if an organism could read genes from an unrelated species. In the 1970s, Stanley Cohen and Herbert Boyer tested this question and found that they were able to move the frog gene for eukaryotic rRNA into E. coli bacteria. These bacteria were able to synthesize both prokaryotic and eukaryotic rRNA making exciting new possibilities in genetics possible.
Complementary base pairing is just one of the important structural features of DNA. In this Activity we will examine many different technologies that manipulate the shape and structure of DNA. Look for these changes as you work through this Activity.
 How is shape important in Biology?
How is shape important in Biology?
Shape 24. How does using complementary base pairing help the enzymes in the Central Dogma copy and read genetic information?
ACTION
Moving DNA
Part of Cohen and Boyer’s experiment involved being able to move frog DNA into bacteria cells. DNA is moved in a structure called a vector. (definition:A vehicle used to transfer the genetic material from the donor organism to the target cell of the recipient organism.) The concept of moving DNA can be applied to almost any cell type. Different vectors are used depending on the size of the DNA being moved, the type of cell receiving the vector, and if the cells are in vivo (definition:comes from Latin for “in life” so it describes living cells.) or in vitro. (definition:comes from Latin for “in glass” so it describes experiments outside of a cell, like in a test tube.) Once a cell has taken in the vector we call it transformed or transgenic.
Let’s explore a few examples.
 Why is shape important in Biology?
Why is shape important in Biology?
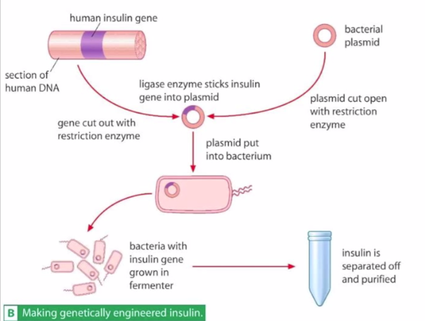
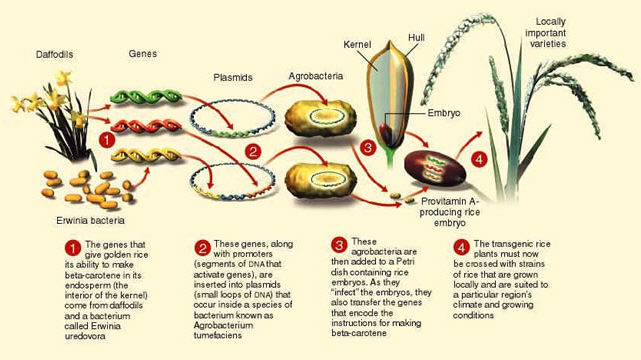
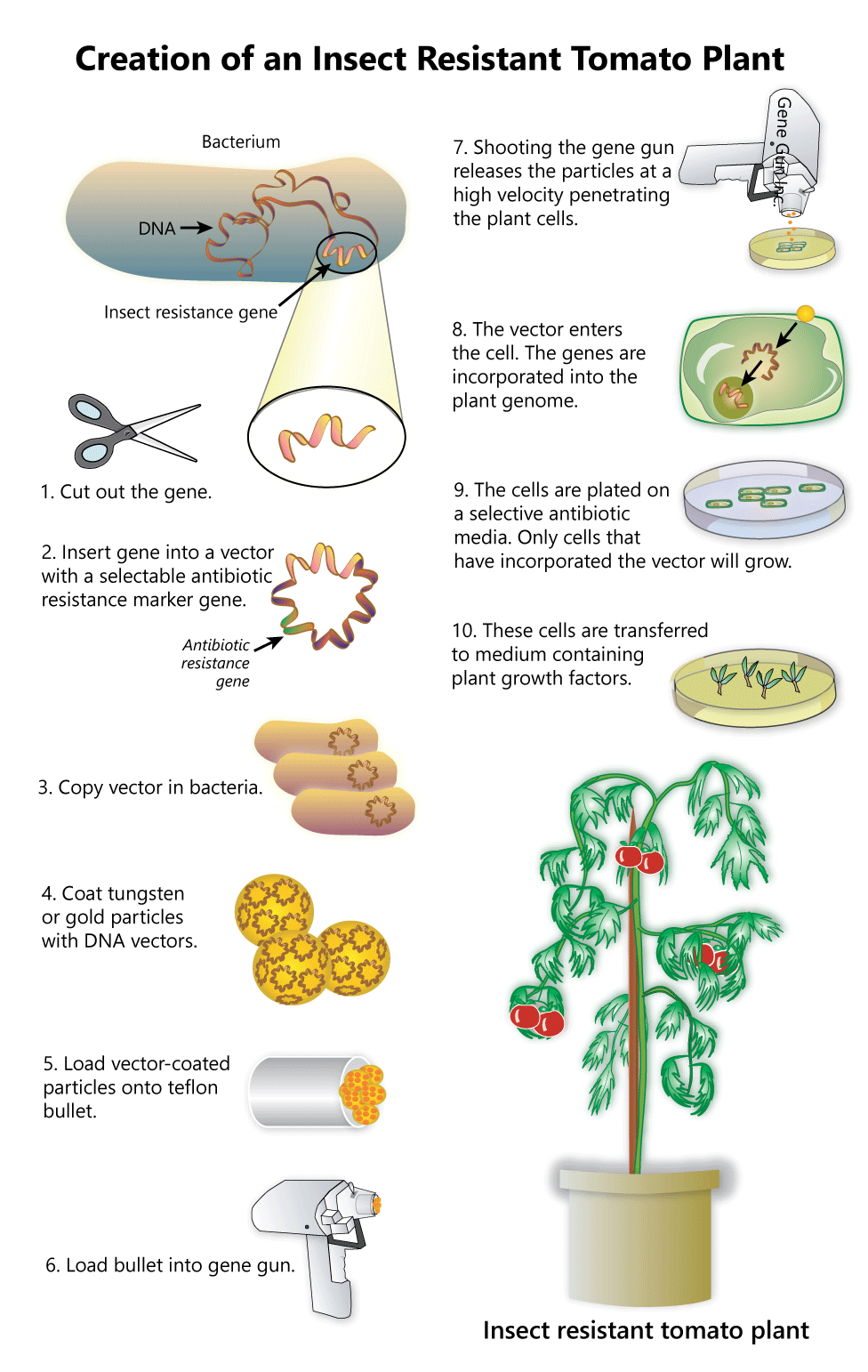
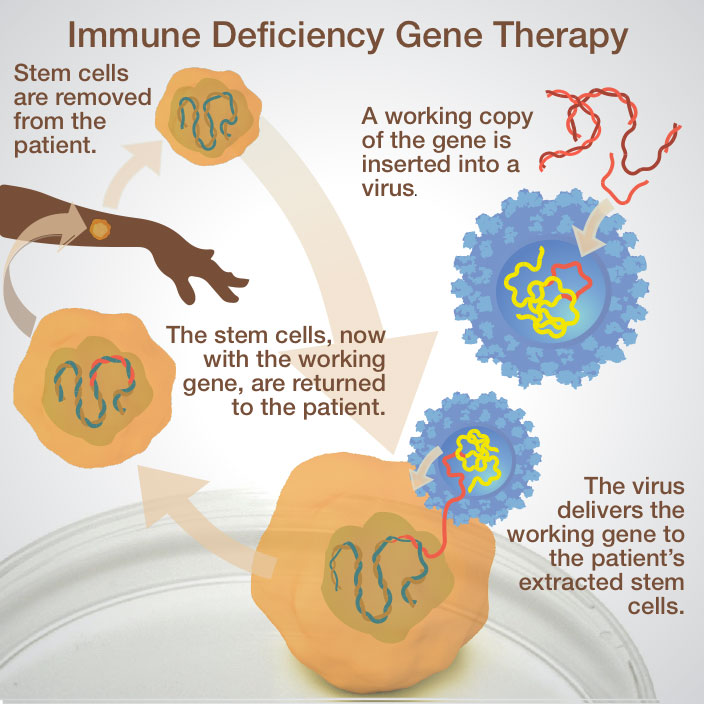
Interpret these pictures showing the steps in moving DNA from one organism to another. Answer these questions for each image:
- What is the source of the DNA?
- What is the target?
- What vector is being used to move the DNA?
- What is the benefit of transforming the target cell in these ways?
Analyzing DNA
In this Activity you will learn about the steps used in different molecular genetics biotechnologies. Your understanding of these steps will help you use good details in this Activity’s tasks.
 Molecular Genetics Biotechnologies
Molecular Genetics Biotechnologies
As you summarize the steps in the biotechnologies described in this Activity look specifically for opportunities to relate how the shape and structure of DNA play a role. The shape can refer to size, length, or nucleotide sequence. Structure can describe the organization of genes on a chromosome, how it’s manipulated, or whether it’s double stranded or single stranded.
Taking DNA and manipulating it requires new methods of working with DNA. For example, different sized pieces of DNA can be separated using a technique called gel electrophoresis. Recall that the phosphate-sugar backbone of DNA is negatively charged because of the phosphate groups. When exposed to electric current DNA will move away from the negative electrode and be attracted by the positive electrode. The mixture of DNA can be forced through a gel made of a web of agarose polysaccharide fibers. This molecular web slows larger fragments of DNA from being able to reach the positive electrode. Look to see how three different sizes of DNA can be separated in this interactive.

Later the DNA can be visualized using dyes that make it visible under UV light. Knowing the size of DNA fragments is a useful tool in many molecular genetics biotechnologies.
For example, gel electrophoresis is used in DNA sequencing. Knowing the exact DNA sequence helps our understanding of how genes are organized on chromosomes. Early work on sequencing DNA involved copying DNA in vitro using many of the components found in cells. Because replication is a complicated process requiring the coordination of many molecules in the right order the process in a test tube was simplified. For example, instead of using primase to make them, primers were made using nucleotide by nucleotide polymerization by simulating the cell’s biochemical reactions. The finished primers can be added to a test tube of DNA and mixed with nucleotides and DNA polymerase to begin replication.
As you watch the following video on sequencing the human genome, look for other differences between replication in sequencing and replication in cells.
The polymerase that is used in this process, Taq polymerase, comes from a thermophilic bacteria. This means that even at high temperatures this enzyme does not denature like those in ordinary bacteria.
One of the challenges in molecular genetics biotechnologies is having and using enough DNA. For example, in the human genome project large amounts of DNA were used to be sequenced. Obtaining a DNA sample from people is simple and painless, usually involving a cheek swab to collect buccal cells from the inside of the mouth. DNA is extracted using a process similar to the investigation you did in Unit 1. However, there are times when the amount of DNA is limited and so it can be copied using the polymerase chain reaction, PCR.
The process in PCR is similar to DNA sequencing in that a sample of DNA, Taq polymerase and primers are introduced into a test tube. Ordinary nucleotides are added as well. Look to see how the steps in PCR differ from those in DNA sequencing in this interactive.
The products can be tested using gel electrophoresis to confirm that the copied DNA fragments are the same size as the original DNA sample.
 How do small steps lead to changes?
How do small steps lead to changes?
Changes 28. Both sequencing and PCR involve parts of reactions in the Central Dogma. Identify the processes that are being recreated in these two biotechnologies. Compare the reactions in these biotechnologies and the processes in the Central Dogma that occur in regular cells.
There are some limitations to PCR, however. Because the reactions cycle for over 50 replications, even small amounts of contamination in the original test tube can be amplified and give false results. Also, Taq polymerase is about as error-prone as other DNA polymerases. Since the DNA polymerase I is not added to this test tube to proofread, any mistakes made in copying DNA can be amplified.
One of the more famous applications of these biotechnologies is in crime scene investigations. DNA samples from a suspect are collected from the crime scene and analysed against samples from a list of suspects. In Canada, the RCMP keeps a database called the DNA Data Bank that includes DNA from unsolved crimes and those from offenders convicted of serious crimes. The following interactive allows you to analyse DNA evidence to help identify a criminal. Before you begin, continue to use good Independent Work skills (definition:I can independently monitor, assess, and revise plans to complete tasks and meet goals.) to review the details in your Portfolio about gel electrophoresis, DNA sequencing, and PCR. The reliability of your conclusions in solving this case depends on your understanding of these processes!

Molecular Toolbox
In the previous Activity we saw how bacteria can be killed by antibiotics. These medicines work by inhibiting structures in prokaryotic cells that are not found in our cells. For example, penicillin inhibits certain bacteria from being able to synthesize the cell wall that protects them from our immune system. Another antibiotic, tetracycline, acts by inhibiting tRNAs from entering the ribosomal A-site preventing protein synthesis. Because prokaryotic ribosomal subunits are smaller than eukaryotic ones, tetracycline only inhibits translation in bacteria.
Antibiotics are a selective force on bacteria. A population of bacteria, like other species, has variations among individuals. Genetic variation by mutations means that it is possible for a mutant bacteria to develop antibiotic resistance. Often the gene for this trait is found on a separate plasmid apart from the rest of the circular chromosome. Among certain bacterial species, plasmids can be transferred from one cell to another. In this way antibiotic resistance can spread in a population.
Viruses are not vulnerable to antibiotics because they lack cellular structures. They carry their genetic information from host cell to host cell. Their protein coat binds to specific molecules on the membrane of their host cell and then they inject their genetic information into the cell. Look back to your graphic organizer from Unit 1, Activity 5 on Nucleic Acids to see that viruses contain both RNA and DNA genomes. Viruses take over the enzymes involved in the Central Dogma in cells. Their promoters attract RNA polymerase more than promoters for their host’s genes. For that reason expression of viral genes increases.
RNA viruses, though, undergo a step before this. They need to convert their RNA into double stranded DNA in a process that goes against the natural flow of information in the Central Dogma. Reverse transcriptase is translated from viral RNA and is then used to reverse transcribe the viral RNA genome into a viral DNA genome. We will see shortly that reverse transcriptase is an incredibly useful enzyme in molecular genetics biotechnology.
In your summary from Unit 2, Activity 9 on the Central Dogma you can see that cells have enzymes that breakdown nucleic acid polymers. We also saw that bacteria can be infected by viruses called bacteriophages. Species of bacteria have enzymes adapted to recognize and destroy bacteriophage DNA called restriction enzymes. These enzymes bind to specific sequences in DNA. Interestingly, these sequences of DNA are 4 to 6 base pairs long and are palindromic. (definition:A word or sequence with the same meaning read forwards or backwards. For example, RACECAR.) They cut the DNA in one of two ways.
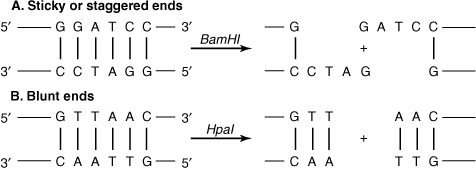
by Clinical Cases Biochemistry for Medics
Blunt ends are made when the double helix is cut so that complementary base pairs are maintained. Sticky ends, on the other hand, are made when single stranded sections of DNA stick out the ends of the cut DNA. They are considered sticky because these exposed nucleotides can reform hydrogen bonds if with the correct sequence of complementary nucleotides.
Restriction enzymes that make sticky ends are particularly useful in molecular genetics biotechnologies. They work like scissors in a molecular-level cut and paste activity. The “glue” that is used to put the pieces together is another enzyme from replication: ligase. Ligase acts by reforming phosphodiester bonds between nucleotides.
These enzymes were all known and understood by the time Cohen and Boyer turned their attention to moving genes from one organism to another. Continue to practice good Independent Work skills (definition:I can independently monitor, assess, and revise plans to complete tasks and meet goals.) by adding details to your Portfolio as you look at the following interactive to see the steps to genetically engineer an organism. Look to see the challenges with the structure of eukaryotic genes that needed to be overcome in this biotechnology.
A genetically engineered chromosome, like a plasmid, with a new gene is sometimes also called a recombinant because it contains a combination of DNA from different sources.
In the interactive we can see that transformed bacteria were screened from the non-transformed bacteria. The transformed bacteria have picked up the plasmid that contains the antibiotic resistance gene allowing them to survive antibiotic treatment. Those bacteria that were not transformed don’t have this trait and are killed by the antibiotic.
 Precise Gene Modifications
Precise Gene Modifications
Recent research in genetic engineering has uncovered an incredibly powerful tool that can be used to modify genes with great precision called clustered regularly interspaced short palindromic repeats, or CRISPR-Cas9. It skips over the steps in making recombinant plasmids and puts it in a vector because the changes are made directly in cells. As you watch this video about CRISPR-Cas9, add information about shape and structures to your Portfolio about Molecular Genetics Biotechnologies.
Medical Applications
Genetic engineering is used to produce a wide variety of hormones and biomolecules. In addition to the example of insulin from the previous interactive, chemical signalling molecules are important in treating other endocrine disorders, blood clotting problems, and cancer are produced in this way. Another possible application involves gene therapy. In this treatment, patients with disease-causing genes receive a normal copy of that gene directly in their cells. The vector used in gene therapy is usually a virus as they are convenient ways of moving genetic information into a patient’s cells.
Currently gene therapy is being used to treat a handful of diseases. Cystic fibrosis is a disease caused by a point mutation in a transporter protein used by cells in many tissues in the body. The transporter is responsible for maintaining chloride ion homeostasis. Learn more about the symptoms of this disorder and then see what makes it a good choice for gene therapy in these two interactives from Genetic Science Learning Center.
In Ontario, babies are screened for the mutation that causes cystic fibrosis. It’s one of 29 diseases that are screened by Newborn Screening Ontario. Provinces across Canada provide a similar service for their newborns. Early detection of these diseases allows for medical choices to help babies receive the best possible care in the years when their bodies are changing the most. On the website the 29 diseases are organized by similar type of disorder. A list of diseases is available under the Disease Information link. For most of these diseases gene therapy has not yet been developed.
At the end of this Activity you will describe biotechnological changes that would need to be done to treat a patient of a disease with gene therapy. Examine one the diseases screened by Newborn Screening Ontario, other than cystic fibrosis. Using other sources, search to find the type of mutation that causes the disorder. This will be the disorder that you will use in your work in the Consolidation task for this Activity.
CONSOLIDATION
Summary
- the structure of DNA can be modified in different ways;
- enzymes can be used to genetically engineer organisms with new traits;
- molecular genetics biotechnologies can be used to treat diseases.
 Using Recombinants
Using Recombinants
You are now ready to complete the assessment for this Activity. You will use the simulation provided to construct a recombinant plasmid used in gene therapy to treat a genetic disorder. The sequence in the cell DNA will simulate the gene for the normal version of the mutated gene for the disorder you are working on. The gene you are looking to change is the same one you identified from Newborn Screening Ontario.
As you prepare for, perform and explain the simulation, look for opportunities to practice good Organization skills. (definition:I can identify, gather and use resources to complete a task.) Also, look for an appropriate format to show how the shape and structure of DNA changes in a clear and organized fashion. Refer back to other ways that you have shown steps in changes earlier in the course. Practice using vocabulary related to molecular genetics biotechnologies to describe the steps in creating this gene therapy vector.
This checklist will help you ensure your work is complete.
Simulation: Using Recombinants
Materials Needed:
- a printout of the three pages with plasmid DNA, cell DNA, and restriction enzymes (using specified colours of paper is optional but helpful.)
- scissors
- tape
Procedure:
- Cut out the PLASMID strips from pink sheet #1. Set aside ANY TWO of the strips (except for the strip which contains the “origin of replication” site (see code at bottom of pink sheet). Shuffle the strips and tape the end of one to the end of another in any random fashion (as long as the letters are going in the same direction). After you have taped the four strips into one long strip, tape the two remaining ends together, to form one long circular paper plasmid. Save the key about antibiotic resistance, from the bottom of the pink sheet for later use. Save extra plasmid pieces until you have successfully completed step #8.
- Cut out the CELL DNA strips from goldenrod sheet #2. They must be taped together in the order indicated at the bottom of each strip. That is, strip 2 is taped to the bottom of strip 1, strip 3 is taped to the bottom of strip two, etc. Note where the DNA code for the normal gene (protein gene) is located.
- It is time to begin testing the various restriction enzymes that you have in your laboratory. Cut out ENZYMES from green sheet #3. There are 8 restriction enzymes given for cutting the DNAs and one ligase for fusing the DNAs together when done. Note that on each of the restriction enzyme rectangles, there is the name of the enzyme (such as Ava II) and a short DNA sequence that shows exactly what sequence that enzyme cuts.
- Use a pen to mark on the PLASMID where each enzyme will cut. Draw the line accurately showing exactly where the bases will be cut apart (and leave the “sticky” ends). Label each. If you have no enzymes that will cut your plasmid only once, then reconstruct your plasmid.
- Mark directly on the CELL DNA strip where the enzymes will cut. Draw the line accurately showing exactly where the bases will be cut apart (and leave the“sticky” ends). Write the name of the enzymes next to each line you draw.
- Your job as a geneticist is to find a restriction enzyme that will cut open your plasmid at ONE site only (this may or may not be possible depending upon how you constructed your plasmid). The same enzyme should be able to cut your cell DNA at TWO sites, one above and one below the normal gene. It is very important that you find an enzyme that cuts as close to this gene as possible. Some of the enzymes cannot cut open your plasmid, some can. Some of the enzymes cannot cut your cell DNA at two sites, some can. If they don’t meet your needs they are not usable.
- After you have completed testing the enzymes, select which ONE enzyme you would use to cut the plasmid and the cell DNA. Use scissors to make the cut in your plasmid and the cell DNA. Be careful to make the cuts in the staggered fashion made by the actual enzyme. This will expose the “sticky” ends where joining will be possible.
- Since one enzyme was used, all “sticky” ends will be compatible. Use tape to splice the normal gene into the plasmid chain. You have created a recombinant plasmid.
- You are interested in confirming that your plasmid contains the normal gene for the disorder you are working on. Describe a test you could use to analyse your DNA.
- The recombinant plasmid is now added to a test tube with the monomers to build the protein coat of viral particles.
- The recombinant virus is now ready to be tested in medical trials to see if it cures patients and if it has any side effects.
If you prefer, you can download a printable copy of this investigation.